|
Name
3-letter code
1-letter code
Neumonic
pka
Relative abundance (%)
Volume (Å^3)
|
Chemical Structure
Click on the images below to view the 3D structures, (i.e., for link to coordinates).
|
|
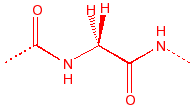
Glycine
Gly
G
Glycine
-
7.5
48
|
flexible, seen in turns
|
|
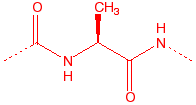
Alanine
Ala
A
Alanine
-
9.0
67
|
hydrophobic, unreactive
|
|
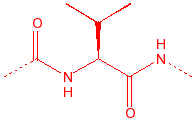
Valine
Val
V
Valine
-
6.9
105
|
hydrophobic, unreactive,
stiff, B-substitution
|
|
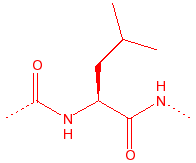
Leucine
Leu
L
Leucine
-
7.5
124
|
hydrophobic, unreactive
|
|
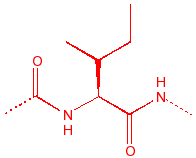
Isoleucine
Ile
I
Isoleucine
-
4.6
124
|
hydrophobic, unreactive,
stiff, B-substitution
|
|
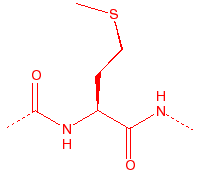
Methionene
Met
M
Methionine
-
1.7
124
|
thio-ether, only unbranched
nonpolar, metal binding site
|
|
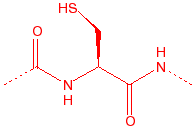
Cysteine
Cys
C
Cysteine
8.33
2.8
86
|
thiol, disulfide
cross-links, binds to iron and zinc, nucleophile in proteases
|
|
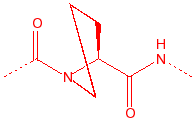
Proline
Pro
P
Proline
-
4.6
90
|
2° amine, stiff,
20% cis, seen in turns
|
|
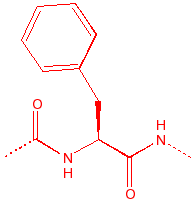
Phenylalanine
Phe
F
Fenylalanine
-
3.5
135
|
hydrophobic, unreactive,
polarizable, UVmax=280nm
|
|
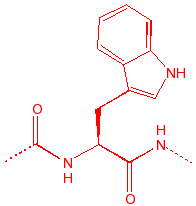
Tryptophan
Trp
W
tWo rings
-
1.1
163
|
largest amino acid, hydrophobic,
fluorescent, UVmax = 250nm, charge transfer
|
|
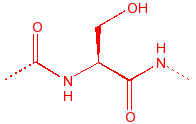
Serine
Ser
S
Serine
-
7.1
73
|
hydroxyl, nucleophile
in serine proteases
|
|
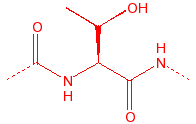
Threonine
Thr
T
Threonine
-
6.0
93
|
hydroxyl, stiff, B-substitution
|
|
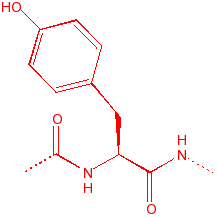
Tyrosine
Tyr
Y
Tyrosine
10.13
3.5
141
|
aromatic, hydroxyl
|
|
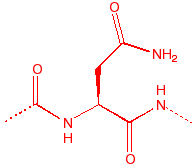
Asparagine
Asn
N
AsparagiNe
-
4.4
91
|
 |
|
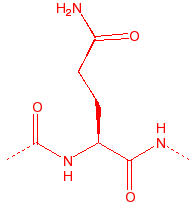
Glutamine
Gln
Q
Qutamine
-
3.9
114
|
 |
|
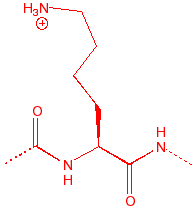
Lysine
Lys
K
Before L
10.79
7.0
135
|
amine base, floppy
|
|
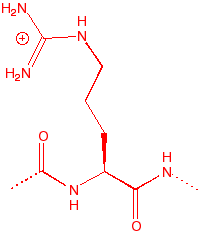
Arginine
Arg
R
Arginine
12.48
4.7
148
|
guanido base, good
charge couple with acid
|
|
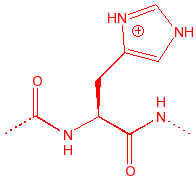
Histidine
His
H
Histidine
6.04
2.1
118
|
imidazole acid or base;
pKa = pH(physio), member of catalytic triad in serine proteases, binds to
copper and zinc
|
|
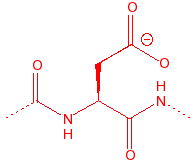
Aspartic Acid
Asp
D
AsparDic
3.90
5.5
91
|
carboxylic acid, in
active sites for cleavage of C-O bonds, member of catalytic triad in serine
proteases, binds to calcium
|
|
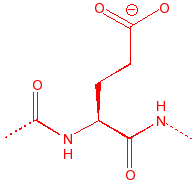
Glutamic Acid
Glu
E
GluEtamic
4.07
6.2
109
|
carboxylic acid, binds
to calcium
|
|
|
|
|
|
|
the end
|
|
|
|
|
|
|
|







































